Introduction
Immunoprecipitation (IP) and coimmunoprecipitation (Co-IP) are widely utilized techniques used for studying protein interactions within complex biological samples by isolating and purifying specific proteins or complexes from them.
Understanding their relationships between one another is vital in deciphering cell processes, signaling pathways and disease mechanisms. Both IP and Co-IP techniques offer unique opportunities to investigate such relationships by selectively isolating proteins of interest or protein complexes from complex samples and purifying them selectively from complex biological samples.
IP and Co-IP techniques rely on antibodies binding specifically with their target antigens in order to capture proteins of interest and their associated interactions. They provide valuable insights into protein composition, dynamics, and functional relationships within a cell context.
In this content outline, we will compare immunoprecipitation and coimmunoprecipitation in detail, outlining their respective principles, applications, advantages, drawbacks, step-by-step procedures involved with each technique as well as any strengths or limitations they might present. Furthermore, we will compare and contrast their objectives, specificity, complexity and analysis of results for comparative purposes.
By understanding these techniques, researchers can select the most appropriate one for their own research goals and gain a greater insight into protein-protein interactions – leading to advancements across multiple areas of biological and medical science.
Definition of Immunoprecipitation
Immunoprecipitation (IP) is an established laboratory technique used to isolate and purify specific proteins or complexes from a mixture. IP utilizes antibody binding sites on antigen targets to selectively precipitate proteins of interest from biological samples along with any of their interaction partners.
Immunoprecipitation works by creating an antibody-antigen complex. Antibodies, known as Immunoglobulins, are produced by our immune systems in Response to foreign substances or antigens which enter our bodies, and these highly specific Proteins bind firmly with target antigens for binding and Recognition.
Immunoprecipitation involves immobilizing antibodies that recognize target proteins on solid supports such as magnetic or agarose beads, before mixing the antibody-coated beads with biological samples containing target and nontarget proteins; selective binding occurs between these antibodies and target protein, creating an immune complex.
To isolate an immune complex from other proteins in a sample, beads are typically washed to remove non-specifically bound proteins before being used to elute out its targeted proteins and associated ones from them. Once isolated, these can then be further studied using techniques like Western blotting, mass spectrometry or functional assays in order to understand their properties and interactions.
Immunoprecipitation is an innovative technique with diverse applications in protein research, from purification and identification of protein modifications to purifying and purifying proteins for further study. Immunoprecipitation allows researchers to study specific roles or behaviors of specific proteins within complex biological systems – providing insights into cellular processes or disease mechanisms.
Definition of Coimmunoprecipitation
Coimmunoprecipitation (Co-IP) is a laboratory technique for studying protein-protein interactions by isolating and identifying those proteins which interact with an unknown target protein in biological samples. Co-IP extends immunoprecipitation by targeting complexes rather than individual proteins as its target.
Coimmunoprecipitation relies on antibodies’ ability to recognize and bind with specific target proteins, leading to precipitation of protein complexes consisting of both them and their binding partners. Like IP, antibodies specific for each target protein are first immobilized onto solid supports like beads or column matrix supports for further study.
Once incubated with antibody-bound support, a sample containing protein mixtures of interest is exposed to antibody-antigen complex formation, with any proteins that interact directly or indirectly with its target being captured and retained by antibody binding sites.
After incubation, the support is washed to remove non-specifically bound proteins and contaminants; only those protein complexes of interest (target protein and its interactants) remain associated with immobilized antibody. Elution methods (competitive elution or use of specific buffers) may then be employed to liberate captured protein complexes from solid support.
These protein complexes can then be further studied using various techniques, including Western blotting, mass spectrometry or functional assays to characterize interacting proteins, identify novel interaction partners and uncover protein-protein interaction networks. Coimmunoprecipitation provides invaluable insights into composition stability dynamics of protein complexes; providing more insight into cellular processes signaling pathways or disease mechanisms.
Coimmunoprecipitation is an invaluable method of protein research, and especially useful in studying interactions between proteins. By capturing and studying protein complexes in their natural environments, researchers gain greater insight into their function within biological systems and how they contribute to functioning properly.
Comparison Table of Immunoprecipitation and Coimmunoprecipitation
Below is a comparison table highlighting the key differences between immunoprecipitation (IP) and coimmunoprecipitation (Co-IP):
| Aspect | Immunoprecipitation (IP) | Coimmunoprecipitation (Co-IP) |
|---|---|---|
| Objective | Isolation and purification of a specific protein | Isolation and identification of protein complexes |
| Targets | Individual protein of interest | Protein complexes involving the target protein and its partners |
| Specificity of Interaction | Detects direct or indirect interactions | Detects predominantly direct interactions |
| Controls and Background Noise | Control samples without the target antibody | Control samples with non-specific IgG antibody |
| Experimental Design | Typically simpler | More complex, involving cross-linking and extensive optimization |
| Sensitivity | High sensitivity due to specific antibody binding | Lower sensitivity due to potential loss of weak interactions |
| Analysis of Results | Individual protein characterization and functions | Protein complex composition, dynamics, and interaction networks |
| Applications | Protein purification, interaction studies | Characterization of protein complexes, pathway analysis |
This table provides a concise overview of the key differences between immunoprecipitation and coimmunoprecipitation.
It highlights the different objectives, target proteins, specificity of interaction detection, controls, experimental design, sensitivity, analysis of results, and applications associated with each technique.
Similarities between immunoprecipitation and coimmunoprecipitation
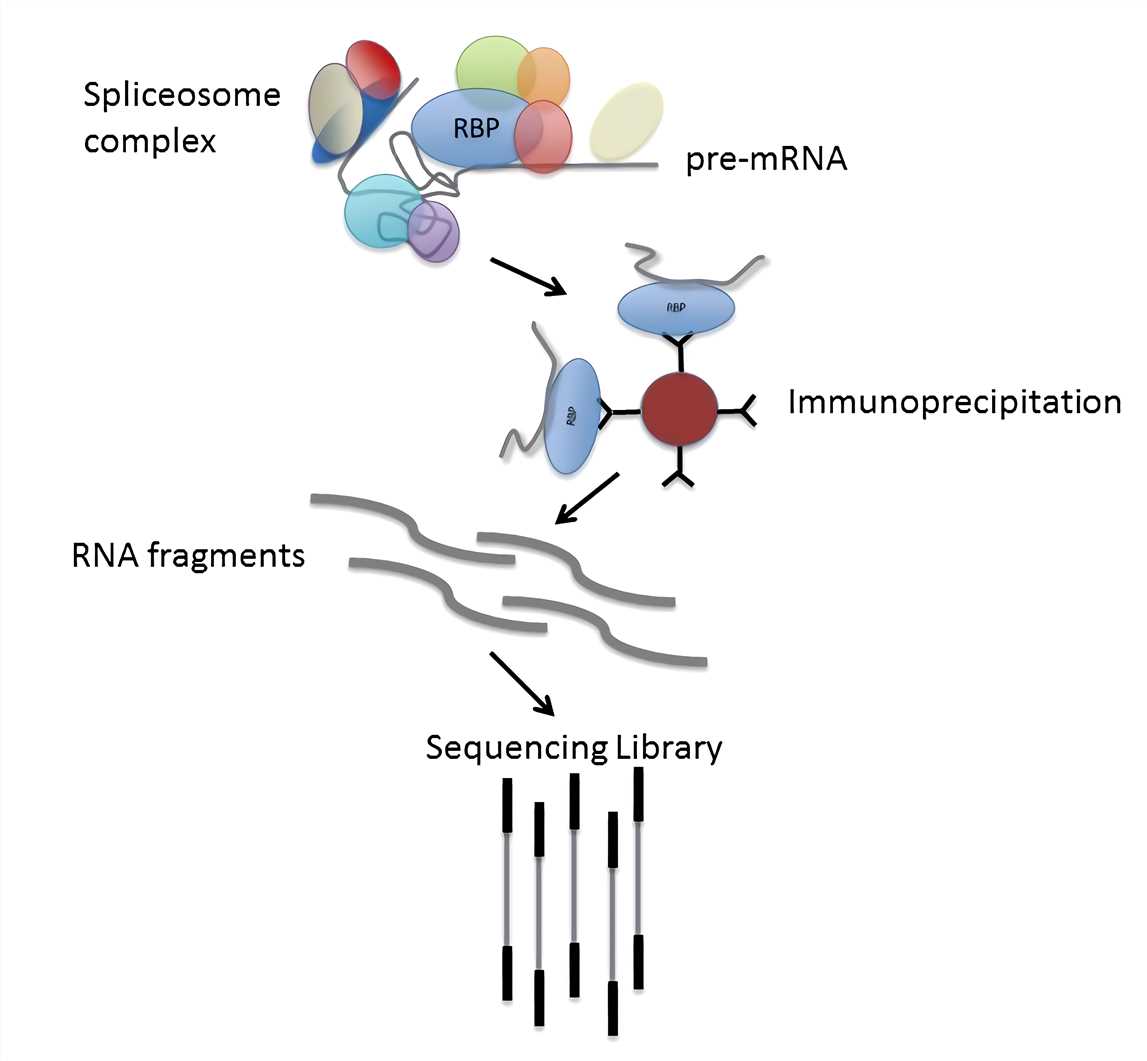
Immunoprecipitation (IP) and coimmunoprecipitation (Co-IP) differ significantly in their objectives and applications, yet both techniques share numerous similarities.
Here are a few key similarities:
Principle of IP and Co-IP: Both approaches rely on antibody-antigen binding as their basis. Specific antibodies used to target proteins or protein complexes allow scientists to capture and precipitate only those of interest in an experiment.
Antibody Selection: Both techniques require carefully selecting and validating antibodies in order to ensure specificity and affinity against target proteins, which will have an enormous effect on the success and reliability of experiments.
Sample Preparation: Both IP and Co-IP involve prepping a biological sample containing proteins of interest by disrupting cells or tissues to release their proteins and create an environment conducive for antibody-antigen binding.
Solid Supports: IP and Co-IP techniques frequently utilize solid supports such as magnetic beads or agarose beads to immobilize antibodies and facilitate their binding to target proteins.
Washing Steps: Both IP and Co-IP require washing steps to remove nonspecifically bound proteins and contaminants, in order to isolate desired proteins or protein complexes while decreasing background noise levels. These washes help isolate desired proteins while simultaneously decreasing background noise levels.
Elution: After precipitation, both IP and Co-IP methods require elution methods to release captured proteins from solid supports for analysis downstream.
Downstream Analysis: Both techniques offer the capability of examining isolated proteins or protein complexes by various means, including Western blotting, mass spectrometry or functional assays. Such analyses provide invaluable information on protein identity, interactions, modifications and functions.
IP is designed to isolate individual proteins while Co-IP allows for the capture of protein complexes and identification of potential partners. Though they differ greatly in methodology and approach, both techniques make for excellent ways of studying protein-protein interactions and their functional implications.
Advantages and limitations of immunoprecipitation
Immunoprecipitation (IP) offers numerous advantages:
Specificity: Immunopharmacetry relies on antibody binding with its target antigen to ensure high specificity for any desired protein, enabling isolation and purification from complex mixtures of proteins of interest.
Versatility: IP can be applied to a broad array of biological samples, including cell lysates, tissue extracts and biological fluids. Furthermore, IP allows researchers to study both soluble and membrane-associated proteins.
Protein Purification: IP is an ideal method for purifying low abundance proteins to obtain highly concentrated samples of them.
Protein-Protein Interaction Studies: IP is an invaluable method for studying protein-protein interactions, precipitating target proteins with their binding partners to identify and characterize protein complexes involved in cell processes.
Protein Modification Analysis: IP can be utilized to examine posttranslational modifications (PTMs) of proteins by immunoprecipitating their modified forms for analysis; such PTMs include phosphorylation, acetylation or ubiquitination.
Limitations of Immunoprecipitation (IP):
Antibody Specificity: IP relies heavily on having access to high-quality antibodies with high specificity and affinity for its target protein, since inappropriate or inferior-quality antibodies could result in nonspecific binding or false positive results.
Loss of Weak Interactions: IP may miss transient or weak protein interactions due to the washing steps used to remove nonspecifically bound proteins; this may result in underrepresentation of certain protein complexes.
Non-Specific Binding: Even when efforts are taken to reduce non-specific binding, IP can still capture proteins that interact indirectly or non-specifically with its target protein, potentially creating background noise and providing background data.
Sample Complexity: In complex biological samples such as tissue extracts or cell lysates, such as tissue extracts or cell lysates, multiple proteins present may prevent efficient isolation and analysis of target protein(s). Pretreatment steps such as fractionation or enrichment may be required in order to reduce sample complexity.
Limited Dynamic Range: IP systems may have limited sensitivity and dynamic range, making it difficult to detect low-abundance proteins or those differentially expressed across samples.
As part of any experiment involving immunoprecipitation, it is crucial to carefully consider its advantages and limitations. Optimizing IP protocol settings, selecting antibodies carefully, and providing proper controls can help mitigate potential restrictions while increasing reliability and accuracy in results.
Advantages and limitations of coimmunoprecipitation
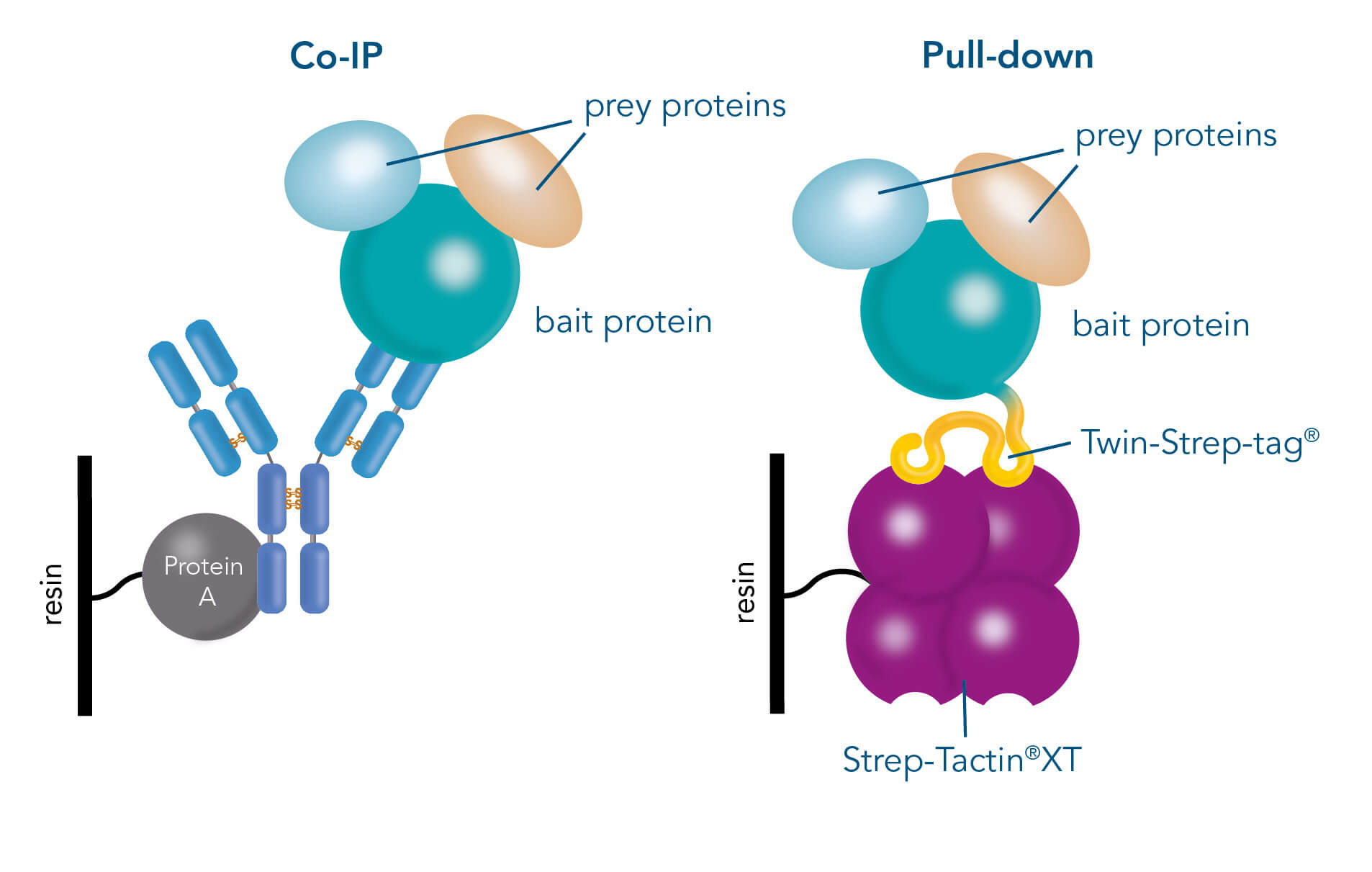
Benefits of Coimmunoprecipitation (Co-IP):
Protein Complex Analysis: Co-IP allows for the capture and study of protein complexes involving target proteins. This offers valuable insights into their composition, dynamics, and functional relationships within a cellular context.
Interaction Specificity: Co-IP allows for the detection of direct protein-protein interactions by precipitating both target protein and its interactant proteins simultaneously, providing a more specific assessment than other techniques.
Novel Interaction Discovery: Co-IP can help identify novel protein-protein interactions, even transient or weak interactions that might otherwise go undetected by other methods. It offers an opportunity to discover previously unknown protein partners and uncover unexpected functional associations.
Validating Protein Interactions: Co-IP can be used to verify protein interactions observed through other techniques, such as yeast two-hybrid or protein-protein interaction databases. By directly recording protein complexes, Co-IP provides direct evidence for physical interactions among proteins.
Pathway Analysis: Co-IP facilitates the characterization of protein-protein interaction networks and signaling pathways, helping elucidate protein roles within complex cellular processes as well as offering insight into disease mechanisms.
Limitations of Coimmunoprecipitation (Co-IP):
Cross-Reactivity and Specificity: As with IP, Co-IP relies heavily on the quality and specificity of antibodies. Cross-reactivity or nonspecific binding of antibodies could result in false-positive results and detection of non-relevant protein interactions.
Complex Experimental Design: Co-IP experiments typically require more involved experimental procedures compared to IP. Cross-linking or intensive optimization may be required in order to stabilize protein complexes and increase specificity.
Sensitivity: Co-IP may have lower sensitivity compared to IP for detecting weak or low-abundance protein interactions, due to extensive washing steps which may wash away weak interactions that would allow detection of transient or less stable complexes.
Co-IP can still capture non-specifically bound proteins or contaminants, leading to background noise and false-positive results. Therefore, non-specific antibodies, such as IgG, must be employed as controls in order to differentiate specific interactions from non-specific binding.
Data Interpretation: Analyzing Co-IP results requires careful interpretation and consideration of potential limitations. Gaining an in-depth knowledge of experimental conditions, protein expression levels and interactions is paramount for accurate data interpretation.
By understanding these advantages and limitations, researchers can optimize Co-IP experiments, select suitable controls, and utilize complementary techniques in order to validate and further explore protein-protein interactions.
Analysis of coimmunoprecipitated complexes
Analysis of coimmunoprecipitated complexes involves several techniques for characterizing their composition, dynamics and functional significance.
Below are some commonly employed for this analysis process:
Western Blotting: Coimmunoprecipitated proteins can be tested using Western blotting to verify the presence of specific proteins within their complex. This technique allows for detection and quantification of individual proteins through specific antibodies.
Mass Spectrometry: Coimmunoprecipitated complexes may be examined using mass spectrometry analysis, which detects and quantifies proteins within them. This approach allows researchers to identify potential interacting partners while providing greater insight into its protein makeup.
Functional Assays: Coimmunoprecipitated complexes can be evaluated using functional assays to measure their biological activity or specific enzymatic functions, including kinase assays, enzyme activity measurements or functional readouts relevant to proteins within the complex.
Immunofluorescence Microscopy: Immunofluorescence microscopy can be used to visualize the localization and co-localization of immunoprecipitated proteins within cells or tissues, providing spatial information regarding interactions partners within these compartments.
Proteomics and Bioinformatics: Proteomic analysis combined with bioinformatics tools is an ideal method for the systematic examination of coimmunoprecipitated complexes, providing insight into novel interactions, protein-protein interaction networks, functional associations or pathways associated with them, etc. This combined approach can reveal novel interacting partners, uncover functional associations or pathways associated with them and uncover functional associations or pathways related to them.
Antibody Blocking/Competition Experiments: Antibody blocking or competition experiments can provide valuable evidence of specificity for coimmunoprecipitation results. By pre-incubating with excess antigen or competing antibodies, binding between target proteins and their interactions partners may be disrupted, providing more evidence for their specificity.
Researchers should employ multiple complementary techniques in order to gain a comprehensive understanding of coimmunoprecipitated complexes. By employing various analytical approaches, they can validate interactions, reveal complex composition and role profiles and gain insights into biological significance of captured protein complexes.
Studying protein complexes and signaling pathways
Studies of protein complexes and signaling pathways are an integral component of understanding various cellular processes and diseases, so understanding them requires engaging in some key techniques and approaches commonly utilized in this research area:
Coimmunoprecipitation (Co-IP): Co-IP is a popular technique to identify protein complexes. By immunoprecipitating target proteins together with their binding partners, this approach facilitates studying protein interactions within signaling pathways and the composition of protein complexes involved in specific signaling pathways.
Mass Spectrometry (MS): MS is a highly accurate method for proteomic analysis, providing precise identification and quantification of proteins within samples. Furthermore, MS can also be used to analyze coimmunoprecipitated complexes to gain insight into their composition, modifications, dynamics and more.
Immunofluorescence Microscopy: Immunofluorescence microscopy allows researchers to observe protein localization within cells and tissues by employing specific antibodies labeled with fluorophores. Researchers can study how proteins distribute themselves within signaling pathways or assess their subcellular localization using this technique.
Western Blotting: Western blotting is a widely employed protein analysis technique that is frequently employed to assess expression levels, post-translational modifications, protein-protein interactions within signaling pathways, as well as probed presence/abundance/interactions of specific proteins of interest. Through specific antibodies Western blotting provides important information regarding presence/abundance/interactions of those proteins of interest.
Functional Assays: Functional assays evaluate the activity or biological function of proteins within signaling pathways, such as kinase activity assays, enzyme assays, reporter gene assays or cell-based assays that measure specific cellular responses triggered by signaling pathway activation.
RNA Interference (RNAi): RNAi is a technique that selectively silences specific genes within cells. By silencing key components of signaling pathways, researchers can explore their impact on overall activity levels and downstream signaling events.
Bioinformatics and Systems Biology: Computational approaches like bioinformatics and systems biology play an indispensable role in studying protein complexes and signaling pathways. By integrating multiple data types – genomics, proteomics, interaction networks – together into holistic understandings of these signaling pathways, regulatory mechanisms identified therein, as well as novel pathway components.
By applying these techniques and approaches, researchers can better understand the composition, dynamics, and functional significance of protein complexes as well as decipher the complex signaling pathways governing cell processes – providing invaluable insight into normal physiology as well as disease mechanisms.
Validation of protein-protein interactions observed by other methods
Validating protein-protein interactions observed via other means is critical in order to ensure accuracy and reliability in findings.
Here are some strategies used for validating such interactions:
Coimmunoprecipitation (Co-IP): Co-IP can be an invaluable way to verify protein-protein interactions detected using other techniques. By immunoprecipitating both target protein and putative interacting partner simultaneously, this technique provides direct evidence of physical association between them.
Pull-Down Assays: Pull-down assays involve immobilizing bait proteins on solid supports such as beads or columns and incubating with protein lysate to capture its interaction partners and isolate them for analysis through Western blotting or mass spectrometry techniques to verify these interactions.
Bioluminescence Resonance Energy Transfer (BRET) and Fluorescence Resonance Energy Transfer (FRET): BRET and FRET are techniques designed to measure protein proximity and interaction using energy transfer between donor and acceptor fluorophores. These tests can also be used to validate protein interactions by adding fluorophores to interact with proteins and measuring the energy transfer signal produced during this process.
Split-Reporter Systems: Split-reporter systems involve breaking apart reporter proteins such as green fluorescent protein (GFP) or luciferase into two nonfunctional fragments that are fused to two proteins of interest, such as GFP or luciferase, before reconstituting into functional reporter protein fragments through protein interaction; these interactions then allow detection of signal using split-reporter systems providing direct readout of protein-protein interactions.
Microscale Thermophoresis (MST): MST is a biophysical technique that measures molecular movement across a temperature gradient. By labeling proteins that interact using various fluorescent dyes, MST can quantify binding affinity as well as confirm interactions based on changes in thermophoretic movement.
Genetic Approaches: Genetic approaches such as yeast two-hybrid (Y2H) or mammalian two-hybrid (M2H) systems can be employed to validate protein-protein interactions by activating or reconstituting reporter genes in response to an interaction between bait and prey proteins.
Rescue Assays: Rescue assays involve functional rescue of phenotypes caused by disruption of protein-protein interactions, with researchers using either their original form or mutant forms of these interactions as a test to see if their rescue phenotype depends on them and provides evidence for the validity of an interaction.
Multiple validation techniques should be utilized to confirm protein-protein interactions identified through other means. Utilizing orthogonal approaches increases reliability of findings and builds confidence in identified interactions.
Sensitivity and specificity of interaction detection
Sensitivity and specificity are key metrics used to measure the performance of interaction detection methods, specifically protein-protein interaction detection. Here’s an explanation of these terms in relation to protein-protein interaction detection:
Sensitivity: Sensitivity refers to an interaction detection method’s ability to correctly recognize true positive interactions, or more broadly speaking, the proportion of true interactions detected correctly by it. A high sensitivity indicates that its rate of false negatives is low enough that more real interactions can be detected than false ones.
Specificity: Specificity is a measure of an interaction detection method’s ability to reliably exclude false positive interactions, quantifying how many reported interactions actually represent true positives. A high specificity indicates low rates of false positives and high confidence in detected interactions.
When assessing an interaction detection method, it’s essential to strike a balance between sensitivity and specificity. While increased sensitivity helps capture as many true interactions as possible, it could also increase false positives; while increased specificity reduces likelihood of false positives but could lead to false negatives that prevent capture of some interactions that truly exist.
To assess the sensitivity and specificity of an interaction detection method, various validation approaches may be taken:
Control Experiments: Running control experiments using non-interacting proteins or negative controls is crucial in order to assess both background levels of false positives and specificity of methods used.
Gold Standard Datasets: When assessing interactions, comparison with gold standard datasets containing known interactions is an excellent way of gauging both their sensitivity and specificity. Overlaps and discrepancies between detected interactions and gold standard datasets provide insights into the performance of the method being employed.
Independent Validation: Employing independent validation techniques such as Co-IP or split-Reporter assays to assess selected interactions can provide a holistic evaluation of their interactions and can confirm true positives while eliminating false positives to provide an estimate of method specificity.
Receiver Operating Characteristic (ROC) Analysis: The Receiver Operating Characteristic (ROC) analysis plots the true positive rate (sensitivity) against false positive rate (1 – specificity) across various cutoffs or thresholds used in interaction detection methods to help determine an ideal balance between sensitivity and specificity, providing a quantitative measure of overall method performance.
Researchers can ensure accurate detection methods by considering both sensitivity and specificity, along with using appropriate validation strategies, in order to assess their reliability and accuracy for protein-protein interaction analysis. This provides for robust and meaningful analysis.
Significance of immunoprecipitation and coimmunoprecipitation in protein research
Immunoprecipitation (IP) and coimmunoprecipitation (Co-IP) are indispensable techniques in protein research, playing an invaluable role in investigating protein-protein interactions, characterizing complexes formed from proteins, and studying function of proteins. Here are the main uses for IP and Co-IP:
Protein-Protein Interaction Analysis: IP and Co-IP techniques are widely utilized to investigate protein-protein interactions. By selectively precipitating a target protein or complex of interest, these techniques allow for the identification and characterization of interaction partners. Understanding protein-protein interactions is key in deciphering cellular processes, signaling pathways, and macromolecular complex formation processes.
Complex Composition Determination: Co-IP allows for the capture and analysis of protein complexes associated with a target protein. By isolating these complexes, Co-IP provides insight into their composition and stoichiometry, as well as allowing identification of core components, regulatory subunits, associated proteins as well as unraveling functional roles within protein complexes.
Protein Localization Studies: IP and Co-IP can be combined with immunofluorescence microscopy or subcellular fractionation techniques to study the subcellular localization of proteins or complexes. By capturing specific cellular compartments where proteins may reside, these techniques provide valuable insight into protein localization patterns, organelle targeting strategies, and trafficking routes.
Post-Translational Modification Analysis: IP and Co-IP can be employed to analyze post-translational modifications (PTMs) of proteins. By immunoprecipitating modified proteins or their interacting partners, these techniques allow for identification of PTMs such as phosphorylation, acetylation, ubiquitination or methylation – providing insight into their regulatory or functional consequences in cell processes.
Functional Studies: IP and Co-IP allow for the isolation of proteins or protein complexes involved in specific cellular functions or biological processes, for further characterization through functional assays such as enzymatic activity assays or readouts to investigate their biological activities and roles within signaling pathways.
Drug Target Identification: Intellectual Property and Co-IP are invaluable tools in drug discovery and development. By capturing protein targets involved in disease pathways, these techniques help identify potential drug targets while assessing efficacy of drug candidates. Furthermore, IP/Co-IP techniques may also be used to study mechanisms of action as well as their effects on protein interactions or complexes.
Validation of Protein-Protein Interactions: IP and Co-IP provide vital validation of interactions identified using other techniques, such as yeast two-hybrid screens, protein interaction databases or computational predictions. By verifying physical association of proteins these techniques improve reliability and confidence in identified interactions.
Immunoprecipitation and coimmunoprecipitation techniques are indispensable tools in protein research, providing essential insights into protein function, cellular processes and disease mechanisms – making these essential tools for modern protein studies.
Conclusion
Immunoprecipitation (IP) and coimmunoprecipitation (Co-IP) are powerful techniques in protein research that allow researchers to study protein interactions, identify complexes formed of multiple proteins, and understand protein function. Both techniques have immense value when it comes to understanding cellular processes, signaling pathways, disease mechanisms and potential therapies.
IP and Co-IP share similar principles and techniques, such as antibody-targeted targeting, protein isolation and downstream analysis. Both approaches provide insight into protein interactions as well as helping identify and validate possible interacting partners.